Assistant Professor, dougvh@coastal.edu
Department of Biology
Smith Science Center, 126-C
Conway, SC 29528
| Funding |
NSF. 2013-2015. RUI: Proteasomal removal of selenoproteins in plants. MCB-1244009 NSF. 2010-2013. Co-PI. Collaborative Research: Regulation of CpNifS/CpSufE1-mediated iron-sulfur cluster synthesis in plant plastids. Implications for sulfur and iron metabolism and selenium tolerance. MCB-0950648. |
| Research Interests |
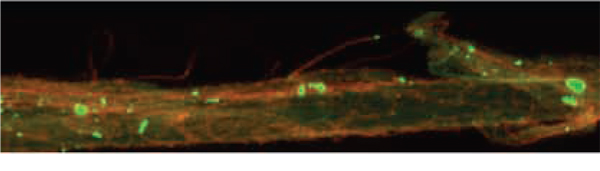
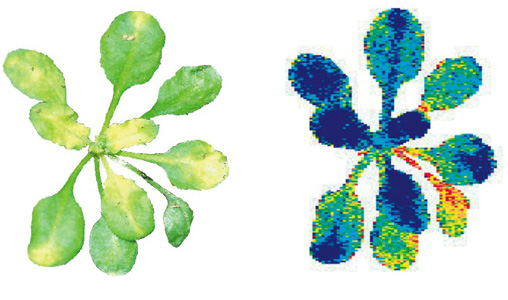
Both natural and human-induced activities have led to contamination of soil and water in the environment. Environmental pollutants can be associated with increased health risks and a decrease in arable land and drinking water. Cleaning polluted lands is of great concern, but is often viewed as an economic burden. Phytoremediation is the use of plants to naturally clean and remediate polluted soil and groundwater, and can be less of an economic strain compared to conventional engineering approaches. Most of my phytoremediation-realted research has focused on understanding pathways that lead to increased plant tolerance of selenium, which at high levels is a public health concern in the Rocky Mountain region. There is currently one project underway that is relavent to phytoremediation, which examines the ability of blackneedle rush- a dominant plant in marsh and estuaries ecosystems in southeastern US- to accumulate excessive amounts of zinc in their tissue. XRF and XANES analyses at the ALS synchotron in Berkeley, CA, suggest that zinc accumulates on the root surface as an inorganic form of hydrated zinc-phoshphate (zinc-hopeite and zinc-hydroxyapatite). Further characterization of these plants is underway, and may reveal insight into the physiology of these plants.
Above: XRF mapping of iron (red) and zinc (green) in roots of blackneedle rush. |
| Regulation and Role of the Proteasome during selnium stress |
Selenium toxicity is thought to occur when selenocysteine replaces cysteine in proteins; this potentially can interfere with disulfide bridges that typically maintain protein structure. Improperly foldeded proteins- ones that may contain diselenide bridges- are removed by the ubiquitin-proteasome system. The proteasome is a large multi-protein complex in eukaryotic cells that is involved in the selected removal of misfolded proteins. On a cellular level, ubiquitin binds to misfolded proteins, which targets the misfolded proteins to the proteasome where they are eventually destroyed by proteases. Therefore, the proteasome prevents cellular stress by preventing the formation of misfolded protein aggregates. My lab has been testing the "malformed selenoprotein hypothesis" by exploring whether or not the proteasome specifically removes misfolded selenoproteins. The role of the proteasome in removing malformed selenoproteins has been explored in photosynthetic organisms with a wide range of selenium tolernance; these plants include the selenium hyperaccumulator Stanleya pinnata, canola, arabidopsis, and the green alga Chlamydomonas. Two central questions govern this project. First, is proteasomal activity correlated to plant species' ability to tolerate selenium or heavy metals such as zinc or cadmium? Secondly, we are exploring the possibility that proteasome activity is regulated by cellular redox state, such as the concentration of glutathione. Selenium stress, as well as other types of stress caused by heavy-metals, is known to induce oxidative stress that can deplete glutathione and cause an imbalance in the redox state. Intriguinly, oxididative stress is also known to inhibit proteasome activity. Higher levels of proteasome activity in plants that are especially tolerant to selenium or cadmium might be maintained by enhanced levels of glutathione. Ultimately, we would like to determine how the proteasome is regulated and possibly maintained during abiotic stress.
|
| Modes of selenium toxicity in plants |
Selenium can cause toxicity in plants becasuse it can be misincorporated into malformed selenoproteins. Additionally, selenate and selenite are pro-oxidants and capable of inducing oxidative stress, which can wreak havoc in cells by oxidizing proteins and lipids. Currently, we are testing whether or not selenite in plant roots inhibits mitochondrial respiration. Specifically, the lab is deciphering if selenite treatment impairs the citric acid cycle by inactivating aconitase and altertering NADH and amino acid metabolism. Additioanlly, the lab is interested in determining if selenium-induced oxidative stress impairs iron-sulfur clusters, which are important cofactors required by many proteins in the electron transport chains.
|